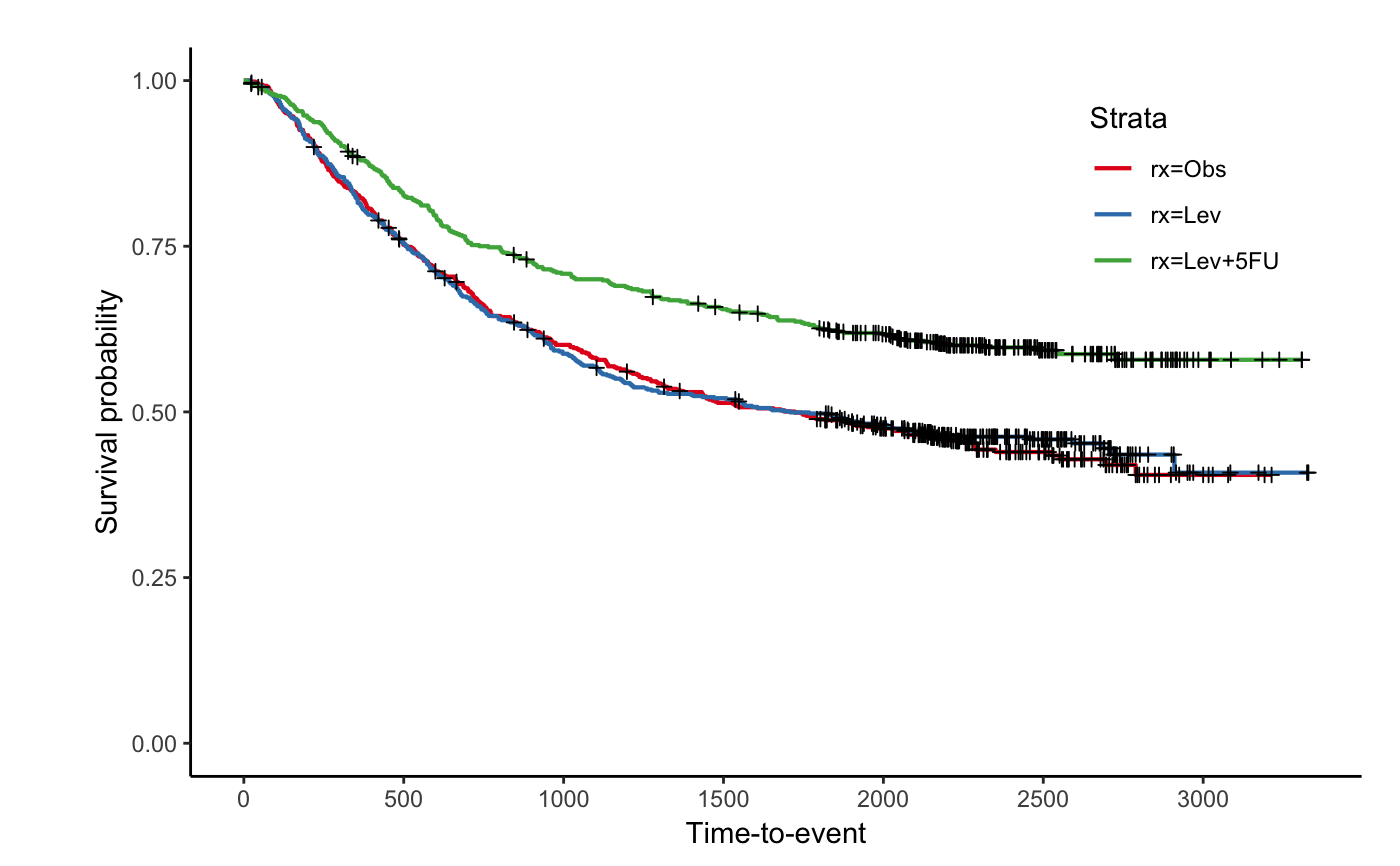
Creates a Kaplan-Meier plot with at risk tables below for survfit object.
Usage
jskm(
sfit,
table = FALSE,
table.censor = FALSE,
xlabs = "Time-to-event",
ylabs = NULL,
xlims = c(0, max(sfit$time)),
ylims = c(0, 1),
surv.scale = c("default", "percent"),
ystratalabs = NULL,
ystrataname = "Strata",
timeby = signif(max(sfit$time)/7, 1),
main = "",
pval = FALSE,
pval.size = 5,
pval.coord = c(NULL, NULL),
pval.testname = T,
marks = TRUE,
shape = 3,
med = FALSE,
legend = TRUE,
legendposition = c(0.85, 0.8),
ci = FALSE,
subs = NULL,
label.nrisk = "Numbers at risk",
size.label.nrisk = 10,
linecols = "Set1",
dashed = FALSE,
cumhaz = F,
cluster.option = "None",
cluster.var = NULL,
data = NULL,
cut.landmark = NULL,
showpercent = F,
status.cmprsk = NULL,
linewidth = 0.75,
theme = NULL,
nejm.infigure.ratiow = 0.6,
nejm.infigure.ratioh = 0.5,
nejm.infigure.xlim = NULL,
nejm.infigure.ylim = c(0, 1),
surv.by = NULL,
nejm.surv.by = NULL,
hr = FALSE,
hr.size = 5,
hr.coord = c(NULL, NULL),
hr.testname = F,
...
)Arguments
- sfit
a survfit object
- table
logical: Create a table graphic below the K-M plot, indicating at-risk numbers?
- table.censor
logical: Add numbers of censored in table graphic
- xlabs
x-axis label
- ylabs
y-axis label
- xlims
numeric: list of min and max for x-axis. Default = c(0,max(sfit$time))
- ylims
numeric: list of min and max for y-axis. Default = c(0,1)
- surv.scale
scale transformation of survival curves. Allowed values are "default" or "percent".
- ystratalabs
character list. A list of names for each strata. Default = names(sfit$strata)
- ystrataname
The legend name. Default = "Strata"
- timeby
numeric: control the granularity along the time-axis; defaults to 7 time-points. Default = signif(max(sfit$time)/7, 1)
- main
plot title
- pval
logical: add the pvalue to the plot?
- pval.size
numeric value specifying the p-value text size. Default is 5.
- pval.coord
numeric vector, of length 2, specifying the x and y coordinates of the p-value. Default values are NULL
- pval.testname
logical: add '(Log-rank)' text to p-value. Default = F
- marks
logical: should censoring marks be added?
- shape
what shape should the censoring marks be, default is a vertical line
- med
should a median line be added to the plot? Default = F
- legend
logical. should a legend be added to the plot?
- legendposition
numeric. x, y position of the legend if plotted. Default=c(0.85,0.8)
- ci
logical. Should confidence intervals be plotted. Default = FALSE
- subs
= NULL,
- label.nrisk
Numbers at risk label. Default = "Numbers at risk"
- size.label.nrisk
Font size of label.nrisk. Default = 10
- linecols
Character or Character vector. Colour imported from ggsci. Default ="Set1", "black" for black with dashed line, character vector for the customization of line colors.
- dashed
logical. Should a variety of linetypes be used to identify lines. Default = FALSE
- cumhaz
Show cumulative incidence function, Default: F
- cluster.option
Cluster option for p value, Option: "None", "cluster", "frailty", Default: "None"
- cluster.var
Cluster variable
- data
select specific data - for reactive input, Default = NULL
- cut.landmark
cut-off for landmark analysis, Default = NULL
- showpercent
Shows the percentages on the right side.
- status.cmprsk
Status value when competing risk analysis, Default = 2nd level of status variable
- linewidth
Line witdh, Default = 0.75
- theme
Theme of the plot, Default = NULL, "nejm" for NEJMOA style, "jama" for JAMA style
- nejm.infigure.ratiow
Ratio of infigure width to total width, Default = 0.6
- nejm.infigure.ratioh
Ratio of infigure height to total height, Default = 0.5
- nejm.infigure.xlim
x-axis limit of infigure, Default = NULL
- nejm.infigure.ylim
y-axis limit of infigure, Default = c(0,1)
- surv.by
breaks unit in y-axis, default = NULL(ggplot default)
- nejm.surv.by
breaks unit in y-axis in nejm figure, default = NULL(ggplot default)
- hr
logical: add the hazard ratio to the plot?
- hr.size
numeric value specifying the HR text size. Default is 5.
- hr.coord
numeric vector, of length 2, specifying the x and y coordinates of the p-value. Default values are NULL
- hr.testname
logical: add '(Log-rank)' text to p-value. Default = F
- ...
PARAM_DESCRIPTION
Author
Jinseob Kim, but heavily modified version of a script created by Michael Way. https://github.com/michaelway/ggkm/ I have packaged this function, added functions to namespace and included a range of new parameters.